Neuro Developmental Encoding (NDE)¶
The Neuro Developmental Encoding (NDE for short) is a vector based indirect representation method to encode the genotype of modular robot bodies used by default in ARIEL.
Source: publication pending
Part 1: Overview¶
The NDEs work by passing the input vectors through a neural network, and outputting 3 adjacency probability matrices of dimensions MxM, where M denotes the number of robot modules we want to generate. An adjacency matrix encodes the probability of a connection between two modules.
The three adjacency matrices are:
The type matrix holds the probabilities of a specific module being of hinge or brick,
The connection matrix holds the probabilities of two modules being connected, for each of the 6 face of the module
The rotation matrix contains the probabilities of a connected module being rotated.
The possible rotation are 8 in increments of 45°.
Note that 2 hinges connected to each other can only be rotated on 90 degree increments
The evolvable/learnable genotype is in the form of a list of vectors as seen in the figure below.
For more information, please read the published paper (pending).
Part 2: Implementation¶
Needed Imports¶
[ ]:
# NDE imports
import mujoco as mj
# Imports for generating numbers
import numpy as np
# Visualising Matrices
from plotting import plot_matrices
from ariel.body_phenotypes.robogen_lite.constructor import (
construct_mjspec_from_graph,
)
from ariel.body_phenotypes.robogen_lite.decoders.hi_prob_decoding import HighProbabilityDecoder
from ariel.ec.genotypes.nde import NeuralDevelopmentalEncoding
# Robot visualisation
from ariel.simulation.environments import SimpleFlatWorld
from ariel.utils.renderers import single_frame_renderer
Initialisation¶
The NDE requires 2 parameters to be initialised.
The number of ARIEL Modules it is encoding for, and
The size of the vectors used in the genotype.
[ ]:
# Define number of modules and gene_size
num_modules = 20
# Default value is 64, but we will use 8 for easier visualisation
gene_size = 8
NDE = NeuralDevelopmentalEncoding(number_of_modules=num_modules, genotype_size=gene_size)
Generate Random Genotype¶
[ ]:
# Random genotype of shape (3, gene_size)
# The scale of the values in the vector can affect the diversity of the generated genotypes.
# You can find a detailed breakdown of the scale values in the `tests/functional/nde.ipynb` file.
scale = 64
example_gene = np.random.normal(loc=0, scale=scale, size=(3, gene_size))
[[-18.581 16.084 90.367 -59.906 -24.356 3.947 -35.631 95.090]
[-42.613 4.969 24.225 -33.693 -81.706 112.593 34.475 -18.033]
[140.321 70.105 -44.490 -24.584 64.774 -16.026 -89.251 96.836]]
Generate matrices¶
[ ]:
matrices = NDE.forward(example_gene)
print(matrices[0].shape)
print(matrices[1].shape)
print(matrices[2].shape)
(20, 4)
(20, 20, 6)
(20, 8)
[ ]:
plot_matrices(matrices)
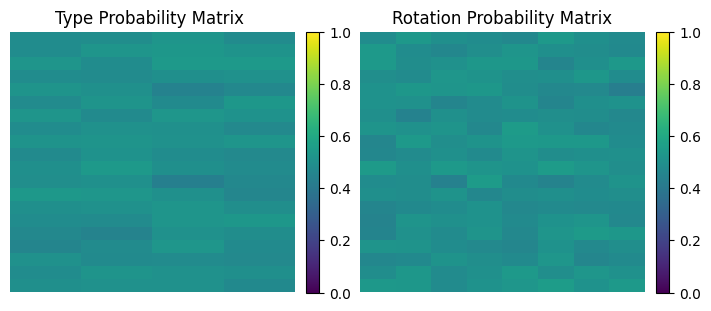
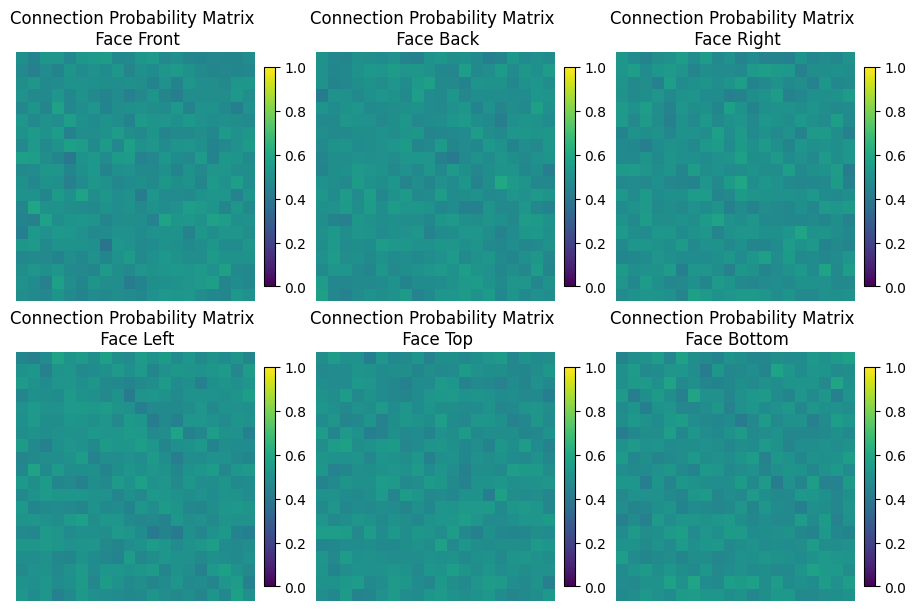
Part 3: Robot Blueprint¶
NDEs were designed to primarily function as a modular robot genotype. The probability matrices don’t really have many other uses (directly). So by default, ARIEL contains functions to turn the NDE genotype into a robot graph.
[ ]:
# Get robot graph from matrices and construct robot specification
def matrices_to_robot(p_matrices: np.ndarray, num_modules: int):
"""Create mujoco specification from robot genotype."""
# Decode the high-probability graph
hpd = HighProbabilityDecoder(num_modules)
robot_graph = hpd.probability_matrices_to_graph(
p_matrices[0],
p_matrices[1],
p_matrices[2],
)
robot_spec = construct_mjspec_from_graph(robot_graph)
return robot_spec.spec
robot_spec = matrices_to_robot(matrices, num_modules)
Spawn robot in simulation¶
Now that we have the robot specification, we can spawn the robot in a world and visualise it.
[ ]:
# Initialise the world you want to spawn the robot in
world = SimpleFlatWorld()
# Spawn the robot
world.spawn(robot_spec)
# Initialise mujoco model and data
model = world.spec.compile()
data = mj.MjData(model)
# Render a frame of the robot
# Zoomed in for better viewing
single_frame_renderer(
model,
data,
cam_fovy=2,
cam_pos=(
-0.7, # x pos
0.1, # y pos
0.5,
), # z pos
)

Evolutionary Operators¶
Since NDE genomes use a vector-based representation, they are compatible with standard evolutionary operators for numeric vectors. Both crossover and mutation can therefore be applied directly to the genome, as long as the structural constraints of the representation are respected.
One important rule is that input vectors must not be mixed during these operations. Although the genome may be stored as a flat vector, parts of it correspond to distinct functional groups and should remain aligned. Evolutionary operators should therefore preserve these boundaries rather than recombining the genome arbitrarily. An example of this is shown in the figure below.
Crossover operators¶
Several crossover operators can be used to recombine two parent genomes into one or more offspring:
Operator |
Description |
|---|---|
|
One-point crossover — selects a single crossover point and swaps the remaining tail segments between two parents. The resulting offspring inherit the first part of the genome from one parent and the second part from the other. |
|
N-point crossover — extends one-point crossover by selecting multiple crossover points and alternating the contributing parent between consecutive segments. This allows more fine-grained recombination across the genome. |
|
Uniform crossover — selects the source parent independently for each gene, usually with equal probability. This produces highly mixed offspring while preserving genome length and overall structure. |
|
Other vector-compatible crossover operators may also be used, provided they preserve the grouping constraints of the representation. |
Mutation operators¶
Mutation operators modify a single genome in-place to introduce variation:
Operator |
Description |
|---|---|
|
Swap mutation — selects two positions in the genome and exchanges their values. This changes the arrangement of genes without introducing new values. |
|
Gaussian mutation — perturbs one or more genes by adding Gaussian noise. This is particularly suitable for real-valued genomes, as it introduces small continuous variations. |
|
Shuffle mutation — selects a subsection of the genome and randomly permutes the values within it. This preserves the same set of values while changing their order. |
|
Other vector-compatible mutation operators may also be used, as long as they do not violate the structural grouping of the genome. |
Representation constraint¶
Although NDE genomes can be evolved using generic vector operators, they should not be treated as completely unstructured arrays. Certain subsets of genes belong together and represent specific input vectors. Mixing values across these boundaries can break the intended semantics of the representation. For this reason, crossover and mutation should always be applied in a way that respects vector boundaries and preserves the internal organisation of the genome.